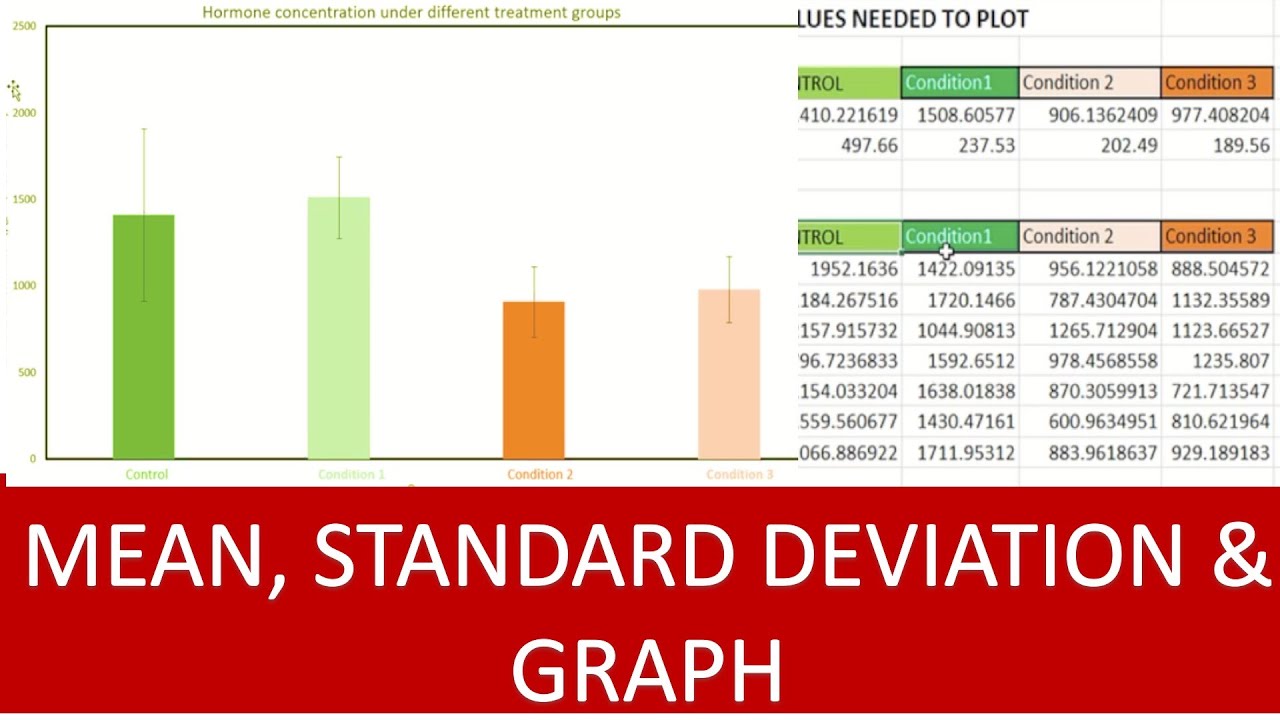
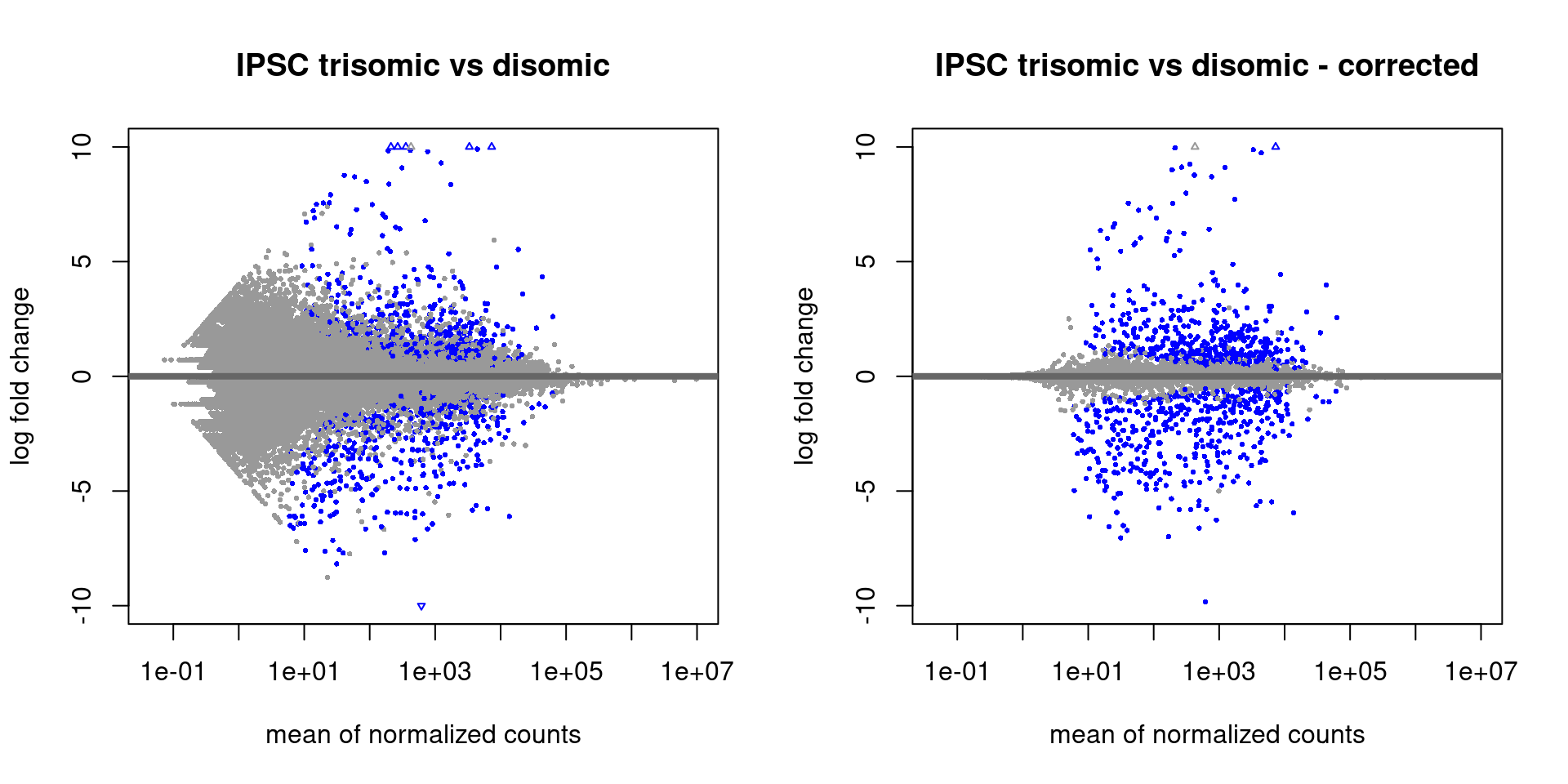
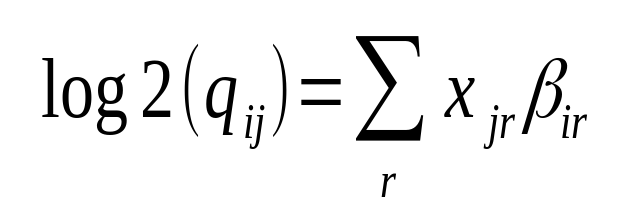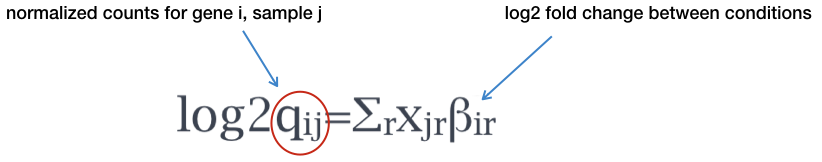Log2 Fold Change Heat Map. A heat map for the 3 different treatments... | Download Scientific Diagram

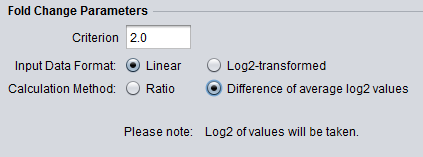
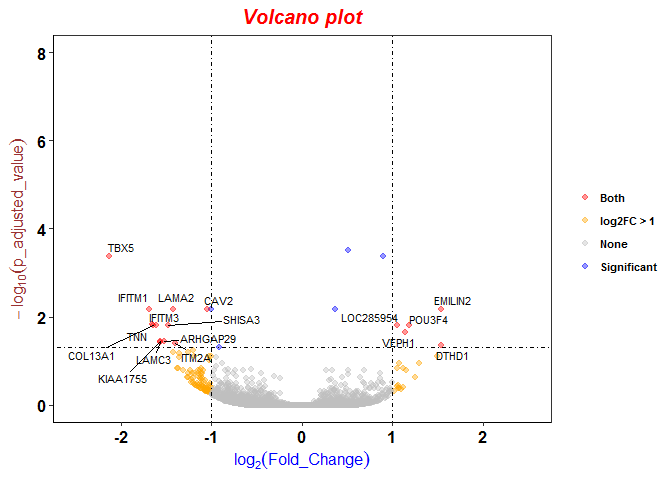
Different logFC (log2foldchange) values for genes from limma-voom and other tools (edgeR and DESeq2)

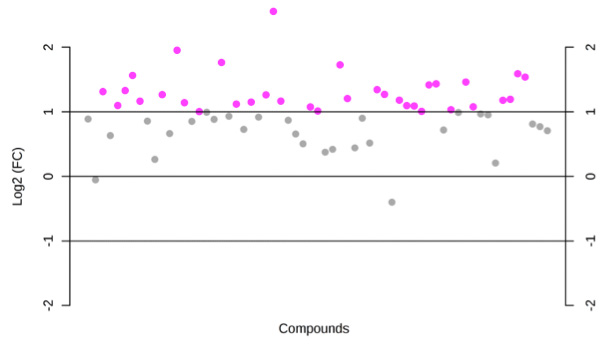
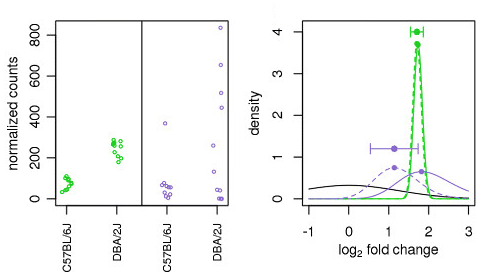
The distribution of log2 fold changes when the expression levels from... | Download Scientific Diagram
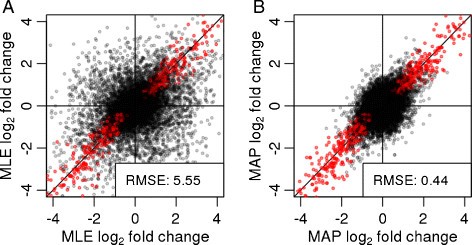
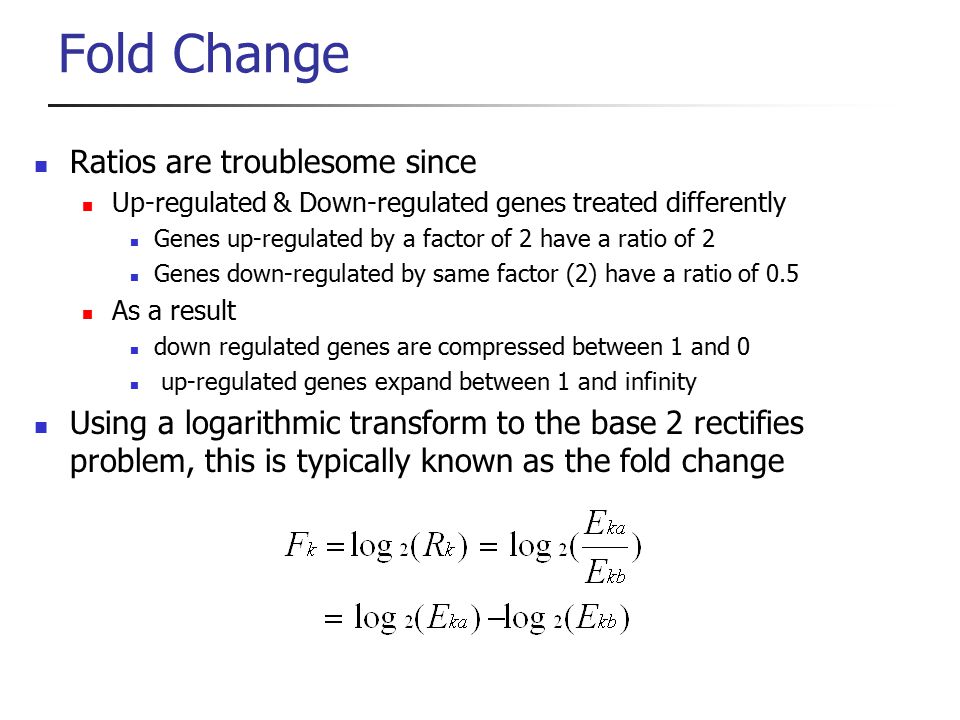
Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2 | Genome Biology | Full Text

Linear regression analysis of fold changes calculated from qPCR and... | Download Scientific Diagram
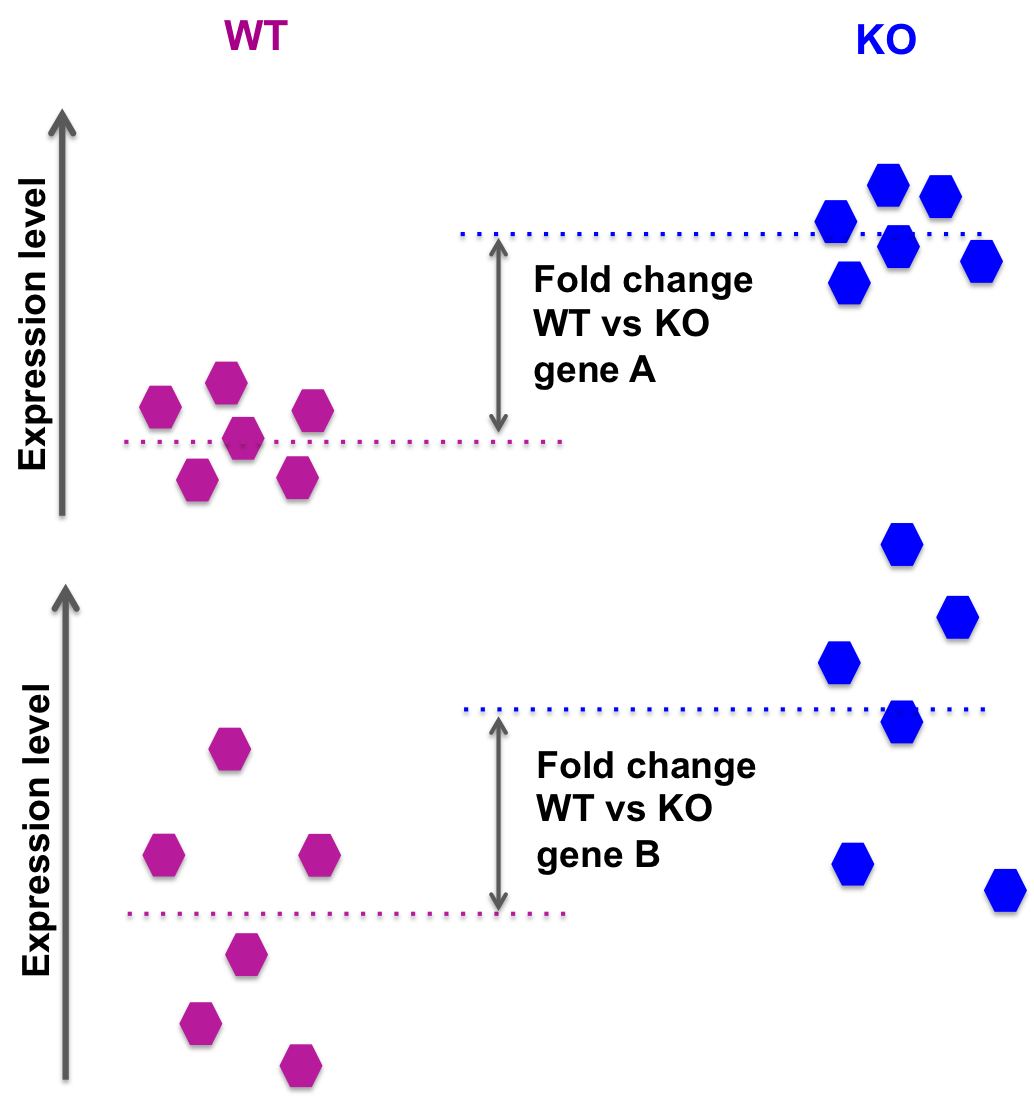
Expression Profiling of Major Histocompatibility and Natural Killer Complex Genes Reveals Candidates for Controlling Risk of Graft versus Host Disease | PLOS ONE
Log2 fold changes in rank_genes_groups are calculated from log-transformed data · Issue #517 · scverse/scanpy · GitHub