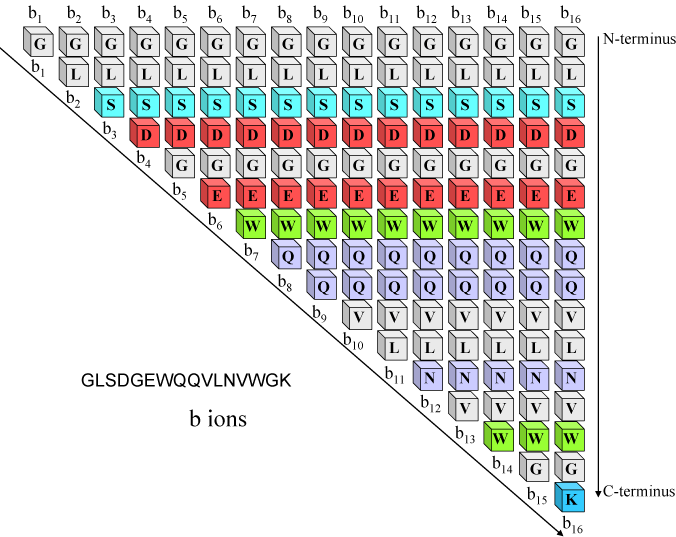
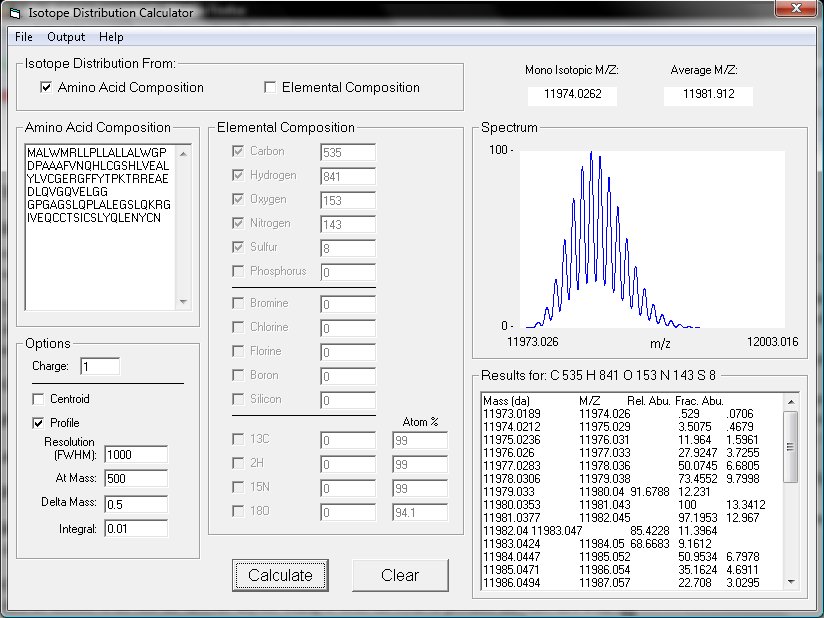
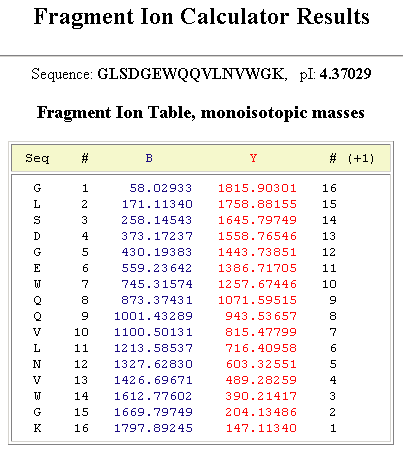
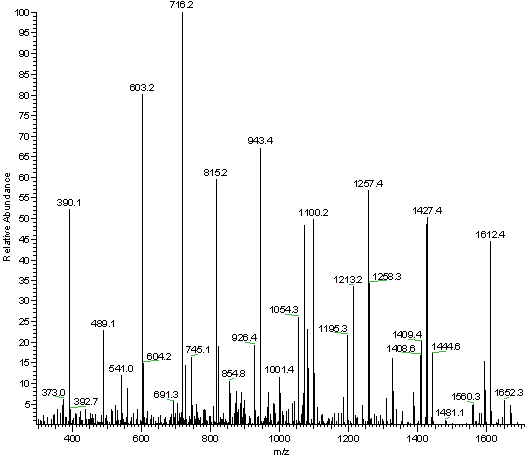
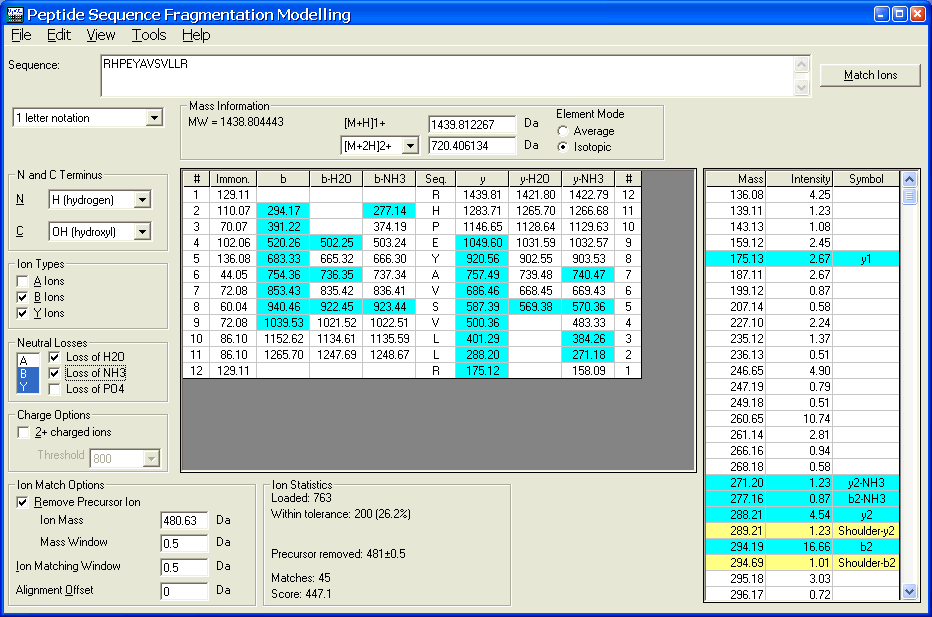
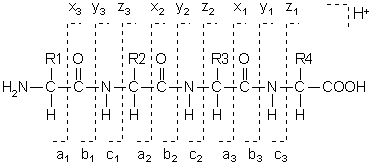The peptide obtained from hydrolysate was identified by ESI-MS/MS mass... | Download Scientific Diagram

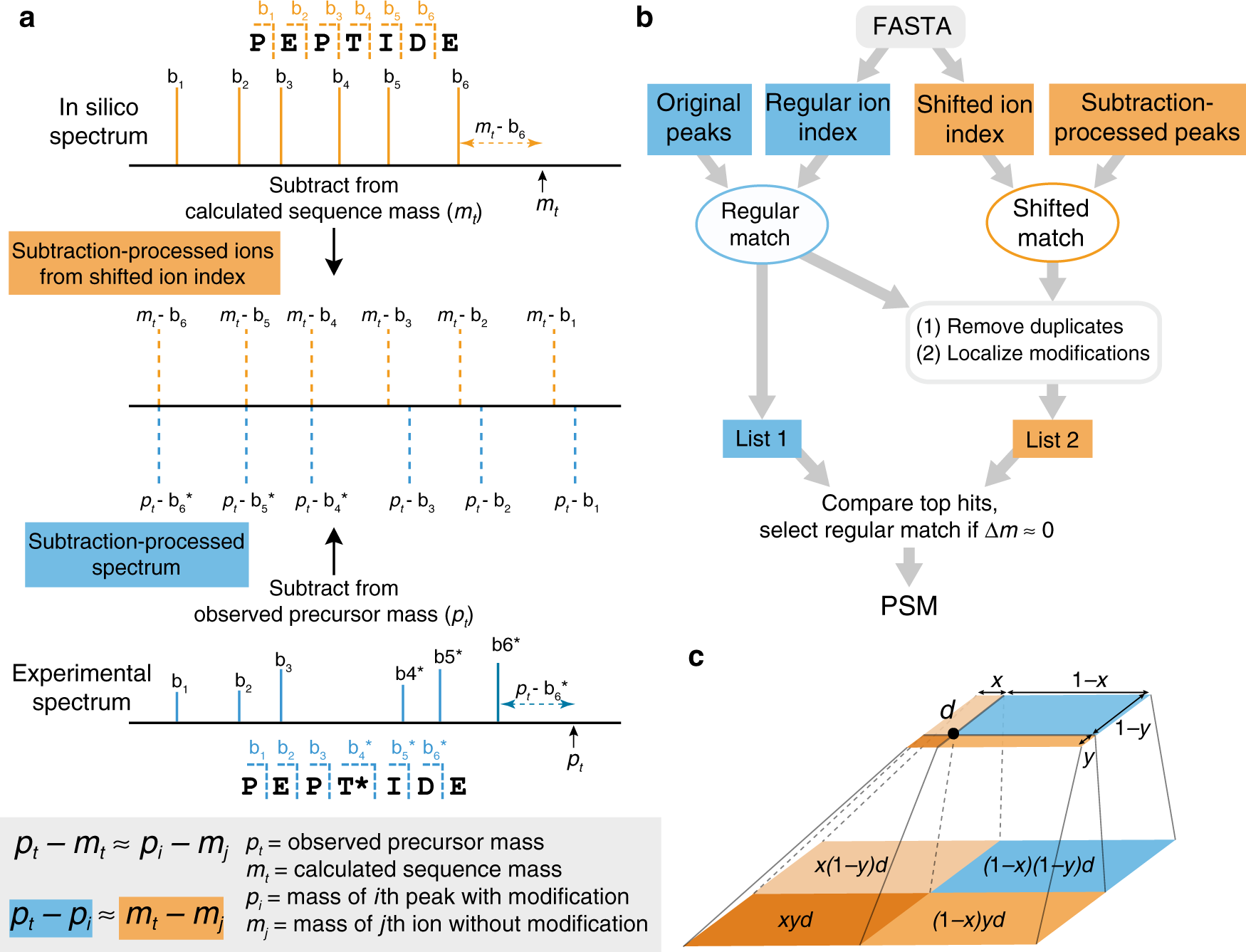
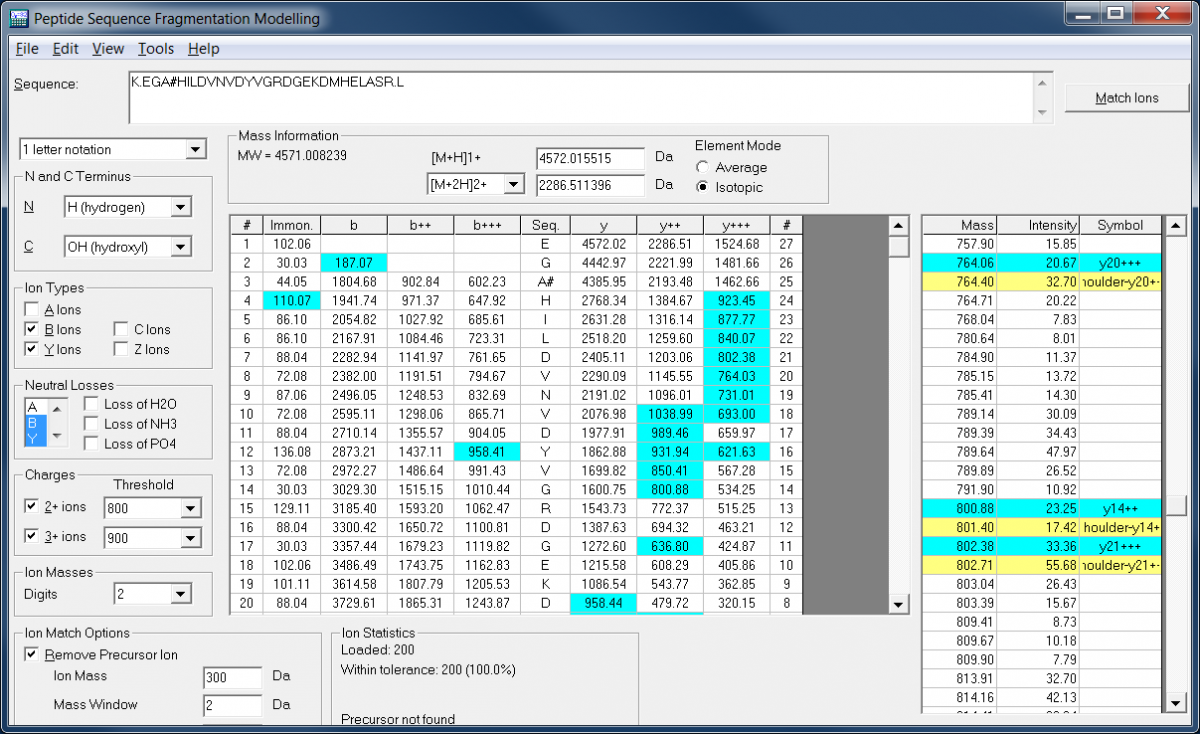
Flowchart summarizing the mass spectral peak assignment algorithm used... | Download Scientific Diagram

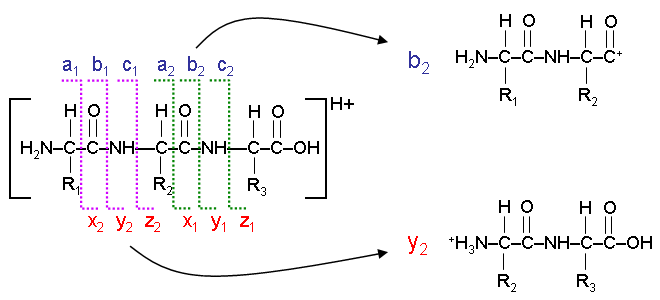
Typical peptide fragmentation generates b or y ions of different mass... | Download Scientific Diagram
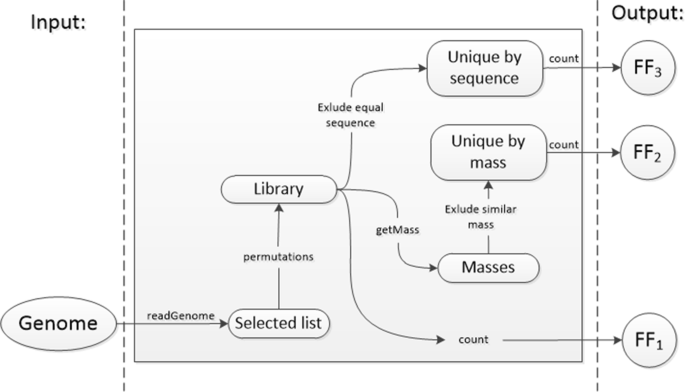
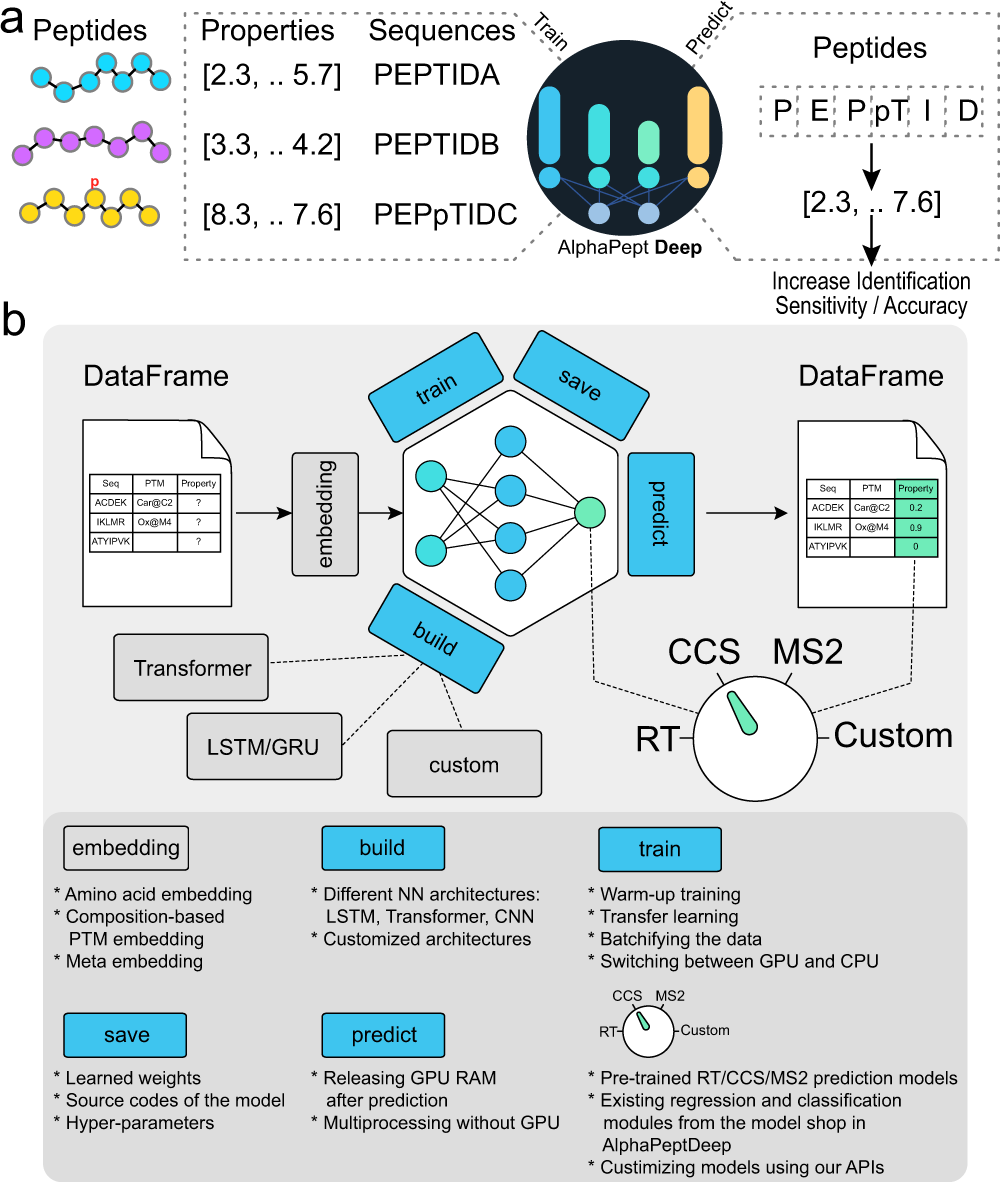
Algorithm-supported, mass and sequence diversity-oriented random peptide library design | Journal of Cheminformatics | Full Text